Conclusions
Introduction
Prostate cancer is one of the most common cancers, with approximately 12.9% of men being diagnosed world-wide. RASSF1A (Ras Associated Domain Family 1A), a tumour suppressor, is associated with the disease and functions to regulate cell growth and apoptosis. Many patients exhibit hypermethylated in the promoter region of RASSF1A, leading to the silencing of the gene. In general, genome wide demethylation is seen with age, yet hypermethylation often occurs in cancer-associated genes. While prostate cancer is rare in men under 40 the risk of developing cancer increases with age.
Systemic demethylation is common in our genomes as we age. However, it is still unknown if the RASSF1A promoter region is targetted for hypermethylation with age.
When choosing a model organism to study I had to make sure it was a mammal as only mammals have prostates. The mouse homolog, RASSF1A haa a very high percentage idendity with the the human protein and contained the same 3 domains. Another reason to use mice is that they are well studied and have preeviously been used in prostate cancer research and show a fimilar disease phenotype.
Prostate cancer is one of the most common cancers, with approximately 12.9% of men being diagnosed world-wide. RASSF1A (Ras Associated Domain Family 1A), a tumour suppressor, is associated with the disease and functions to regulate cell growth and apoptosis. Many patients exhibit hypermethylated in the promoter region of RASSF1A, leading to the silencing of the gene. In general, genome wide demethylation is seen with age, yet hypermethylation often occurs in cancer-associated genes. While prostate cancer is rare in men under 40 the risk of developing cancer increases with age.
Systemic demethylation is common in our genomes as we age. However, it is still unknown if the RASSF1A promoter region is targetted for hypermethylation with age.
When choosing a model organism to study I had to make sure it was a mammal as only mammals have prostates. The mouse homolog, RASSF1A haa a very high percentage idendity with the the human protein and contained the same 3 domains. Another reason to use mice is that they are well studied and have preeviously been used in prostate cancer research and show a fimilar disease phenotype.
Specific Aim 1
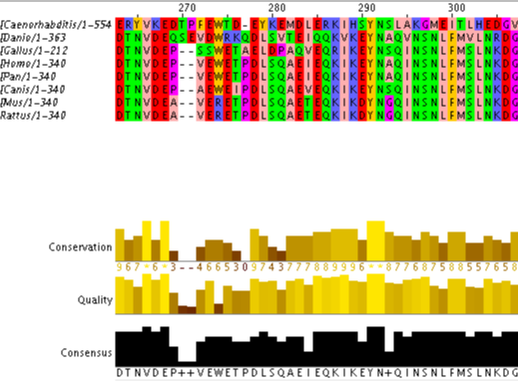
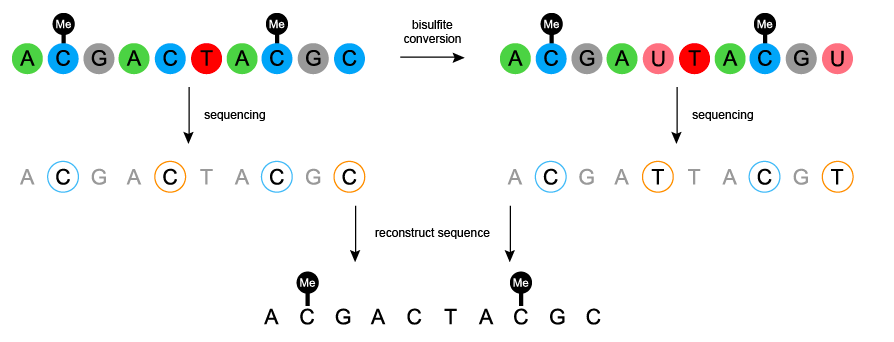
The goal of the first aim is to identify the conservation of the RASSF1A gene and its promoter with various homologs. This will be done by running using Clustal Omega to align the FASTA formatted sequences from each of the RASSF1A homologs, including their promoters. Then to determine each homologs methylation state I will use bisulphite sequencing, each homologs methylation state would be based on the number of remaining cytosine residues as unmethylated cytosine’s would be converted to uracil. I expect to find that the homologs will have very similar conserved regions but the methylation patterns will differ between mammalian species.
The goal of the first aim is to identify the conservation of the RASSF1A gene and its promoter with various homologs. This will be done by running using Clustal Omega to align the FASTA formatted sequences from each of the RASSF1A homologs, including their promoters. Then to determine each homologs methylation state I will use bisulphite sequencing, each homologs methylation state would be based on the number of remaining cytosine residues as unmethylated cytosine’s would be converted to uracil. I expect to find that the homologs will have very similar conserved regions but the methylation patterns will differ between mammalian species.
Specific Aim 2
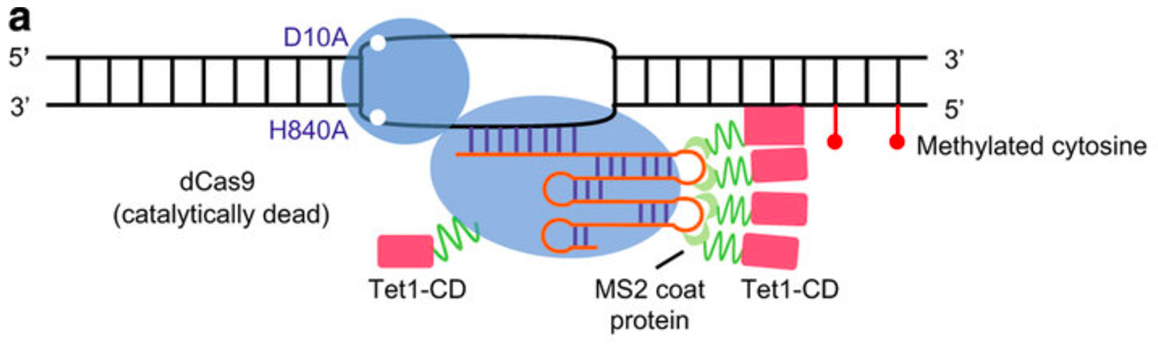
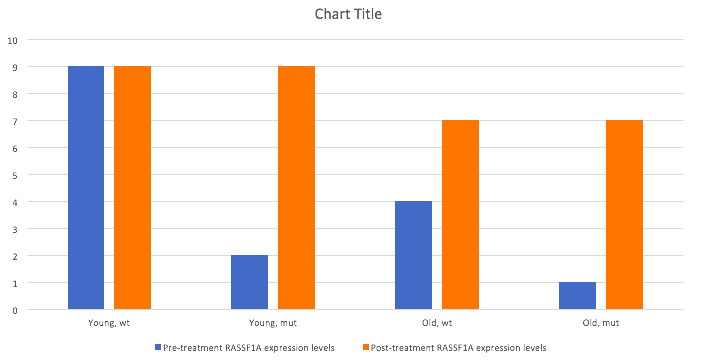
The goal of this aim is to determine conserved methylation sites in the RASSF1A promoter necessary for proper cell growth throughout development. This would be done by creating a CRISPR construct to mutate the conserved methylation sites. The system would be delivered by injection into the prostates of both wt and hypermethylated mutant young and adult male mice. While the CRISPR/Cas9 system will only cause silent mutations and not alter the the amino acid sequence of the protein, the system will carry a Tet1-Cd system which will promote demethylation of the protein. Before and after treatment I would carry out quantitative proteomic analysis using SILAM and mass spectrometry. Altered conserved methylation sites in the RASSF1A promoter will lead to elevated expression, my predicted results can be seen in the graph below, showing a rescued phenotype in both old mice and the young mice with the disease.
The goal of this aim is to determine conserved methylation sites in the RASSF1A promoter necessary for proper cell growth throughout development. This would be done by creating a CRISPR construct to mutate the conserved methylation sites. The system would be delivered by injection into the prostates of both wt and hypermethylated mutant young and adult male mice. While the CRISPR/Cas9 system will only cause silent mutations and not alter the the amino acid sequence of the protein, the system will carry a Tet1-Cd system which will promote demethylation of the protein. Before and after treatment I would carry out quantitative proteomic analysis using SILAM and mass spectrometry. Altered conserved methylation sites in the RASSF1A promoter will lead to elevated expression, my predicted results can be seen in the graph below, showing a rescued phenotype in both old mice and the young mice with the disease.
Specific Aim 3
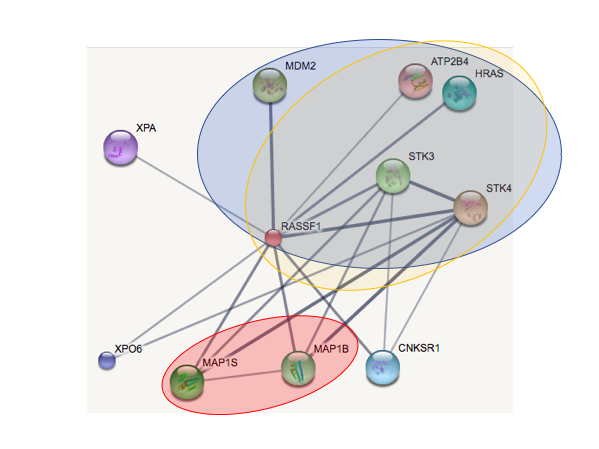
Determine changes in protein interactions after the RASSF1A promoter is hypermethylated. To do this I will isolate protein complexes from prostate cells of WT and mutant hypermethylated mice of different age groups using antibody based co-immunoprecipitation using a bait protein for RASSF1A. I would then carry out liquid chromatography mass spectrometry-based to identify the proteins RASSF1A is interacting with. I will carry out this experiment in young and old mice, with both the wildtype and with the disease. The Ras oncogene proteins are regulated by RASSF1A. With RASSF1A silenced after hypermethylation, I predict it will not be able to function and interact with other proteins as it normally would.
Determine changes in protein interactions after the RASSF1A promoter is hypermethylated. To do this I will isolate protein complexes from prostate cells of WT and mutant hypermethylated mice of different age groups using antibody based co-immunoprecipitation using a bait protein for RASSF1A. I would then carry out liquid chromatography mass spectrometry-based to identify the proteins RASSF1A is interacting with. I will carry out this experiment in young and old mice, with both the wildtype and with the disease. The Ras oncogene proteins are regulated by RASSF1A. With RASSF1A silenced after hypermethylation, I predict it will not be able to function and interact with other proteins as it normally would.
Future Directions
By the end of this study I expect to have a better understanding of the process of increased methylation of the RASSF1 promoter with age and how this hypermethylation affects the resulting proteins interactions. This research will discover more about the changing methylation state of RASSF1A and its promoter during a males aging process. It will also uncover new potential drug targets but identifying protein interactions for the hypermethylated RASSF1A.
By the end of this study I expect to have a better understanding of the process of increased methylation of the RASSF1 promoter with age and how this hypermethylation affects the resulting proteins interactions. This research will discover more about the changing methylation state of RASSF1A and its promoter during a males aging process. It will also uncover new potential drug targets but identifying protein interactions for the hypermethylated RASSF1A.
Final Presentation |
Previous submissions |
|
| ||||||||||||
| |||||||
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.