What are protein domains?
Every protein will have at least one domain within it, which is normally responsible for a function or interaction of the protein and helps the protein do what it is required to do [1]. The same domain can be present in many different proteins, even if the proteins have completely different functions.
What domains are present in RASSF1A?
I carried out my domain analysis for RASSF1A using the FASTA sequence and three different programs available online, SMART, Pfam and Interpro. This allowed me to compare the results from different websites to ensure I was able to identify all the domains present.
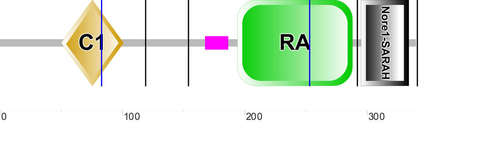
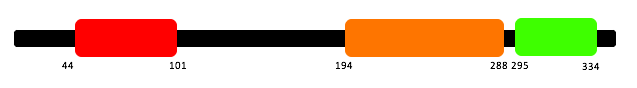
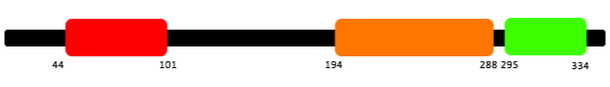
All three programs gave me results showing the same three domains present in RASSF1A: Protein kinase C conserved region 1 (C1), Ras association domain (RA) and Nore1/SARAH (name varies depending on site).
What do these domains do?
The C1 domain is a zinc finger domain which means it is able to make tandem contacts with their target molecules, which in the case of RASSF1A is RasGTP [2]. Studies have shown that the C1 domain is important for death receptor association [3].
The RA domain is the motif that unifies all the Ras family of GTPases. However it is still unclear if the RA domain in RASSF1A is important or if it is able to utilise the Ras to promote cell death. Since RASSF1A only binds weakly to Ras it has been suggested that RA may associate with a different GTPase rather than Ras [4].
The Nore1/SARAH domain is associated with three different families of tumour suppressors (Rassf/Sav/Hpo). The SARAH domain is often associated with C1 domains and RA domains. [5]
What do these domains do?
The C1 domain is a zinc finger domain which means it is able to make tandem contacts with their target molecules, which in the case of RASSF1A is RasGTP [2]. Studies have shown that the C1 domain is important for death receptor association [3].
The RA domain is the motif that unifies all the Ras family of GTPases. However it is still unclear if the RA domain in RASSF1A is important or if it is able to utilise the Ras to promote cell death. Since RASSF1A only binds weakly to Ras it has been suggested that RA may associate with a different GTPase rather than Ras [4].
The Nore1/SARAH domain is associated with three different families of tumour suppressors (Rassf/Sav/Hpo). The SARAH domain is often associated with C1 domains and RA domains. [5]
How well conserved are the domain?
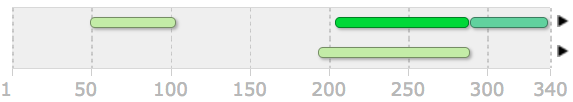
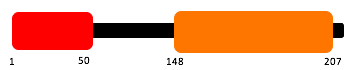
I carried out further domain analysis for the various homologs I identified previously inputting their FASTA sequences into the same three programs. Below shows the results for the 7 homlogs and the amino acid positions for each domains position. Every homolog contains the same 3 domains as the human except for the the chicken which is missing the SARAH domain.
References
[1] EMBL-EBI What are protein domains? https://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification/what-are-protein-domains
[2] SMART C1 domain http://smart.embl-heidelberg.de/smart/do_annotation.pl?DOMAIN=C1&START=50&END=101&E_VALUE=6.64412256768045e-07&TYPE=SMART&BLAST=RGHRFQPAGPATHTWCDLCGDFIWGVVRKGLQCAHCKFTCHYRCRALVCLDC
[3] Foley C.J., Freedman H., Choo S.L., Onyskiw C., Fu N.Y., Yu V.C., Tuszynski J., Pratt J.C., Baksh S. 2008. Dynamics of rassf1a/moap-1 association with death receptors. Mol. Cell. Biol. 28:4520–4535.
[4] Gordon, M. and Baksh, S. 2011. RASSF1A: Not a prototypical Ras effector. Small GTPases. 2(3): 148–157.
[5] Interpro SARAH domain http://www.ebi.ac.uk/interpro/entry/IPR011524
[2] SMART C1 domain http://smart.embl-heidelberg.de/smart/do_annotation.pl?DOMAIN=C1&START=50&END=101&E_VALUE=6.64412256768045e-07&TYPE=SMART&BLAST=RGHRFQPAGPATHTWCDLCGDFIWGVVRKGLQCAHCKFTCHYRCRALVCLDC
[3] Foley C.J., Freedman H., Choo S.L., Onyskiw C., Fu N.Y., Yu V.C., Tuszynski J., Pratt J.C., Baksh S. 2008. Dynamics of rassf1a/moap-1 association with death receptors. Mol. Cell. Biol. 28:4520–4535.
[4] Gordon, M. and Baksh, S. 2011. RASSF1A: Not a prototypical Ras effector. Small GTPases. 2(3): 148–157.
[5] Interpro SARAH domain http://www.ebi.ac.uk/interpro/entry/IPR011524